Lab 9: Future Directions in GIS and Public Health
“Remember: every GPS point is a step toward understanding the world a little better.”
In this final lab, we bring together everything we’ve learned this
quarter:
vector data, raster data, coordinate systems, interactive maps, and
exposure science. In this lab, we’re diving into the wonderful world of
GPS data, our old friend NDVI rasters,
and some fun interactive mapping – with a light touch
and an emphasis on public health applications. This will be our last
interactive session, so we will lean on old tools we’ve been using, and
we will also apply some new packages for the first time!
The objectives of this guide are to teach you to:
- Understand how to load and preprocess GPS tracking data in R
- Calculate movement metrics (distance + speed)
- Create interactive mobility maps with leaflet
- Compare spatial patterns across time of day using
tmap
- Integrate NDVI rasters as environmental exposure
- Extract momentary exposure values
- Visualize exposure over time
Let’s do this, one last time!
Load our packages
library(dplyr) # data wrangling, piping, and general awesomeness
library(readr) # reading in data like a pro
library(terra) # handling raster data
library(sf) # spatial vector data with all the bells and whistles
library(tmap) # beautiful thematic maps, both static and interactive
#New Packages this week!
library(lubridate) # working with dates and times like it's no big deal
library(leaflet) # making interactive maps that aim to impress
library(geosphere) # helps us to colculate the great-circle distance between two points
Import and Prepare Our GPS Data
Here we read in our GPS dataset, parse the timestamp, and create friendly time labels. This is where readr, dplyr, and lubridate come together to easily tackle this task.
download.file(
"https://raw.githubusercontent.com/pjames-ucdavis/SPH215/refs/heads/main/gps_apr25_30.csv",
destfile = "gps_apr25_30.csv",
mode = "wb"
)
gps_data_raw <- read_csv("gps_apr25_30.csv")
gps_data <- gps_data_raw %>%
mutate(
time = ymd_hms(`UTC time`),
time_label = format(time, "%Y-%m-%d %H:%M:%S")
) %>%
arrange(time)
glimpse(gps_data)## Rows: 21,886
## Columns: 8
## $ timestamp <dbl> 1.61931e+12, 1.61931e+12, 1.61931e+12, 1.61931e+12, 1.61931…
## $ `UTC time` <dttm> 2021-04-25 00:15:10, 2021-04-25 00:15:10, 2021-04-25 00:16…
## $ latitude <dbl> 42.37178, 42.37177, 42.37180, 42.37183, 42.37186, 42.37183,…
## $ longitude <dbl> -71.08780, -71.08793, -71.08796, -71.08791, -71.08791, -71.…
## $ altitude <dbl> 16.142262, 3.940258, 3.940258, 3.940258, 3.940258, 3.940258…
## $ accuracy <dbl> 25.62929, 65.00000, 1030.74153, 65.00000, 65.00000, 65.0000…
## $ time <dttm> 2021-04-25 00:15:10, 2021-04-25 00:15:10, 2021-04-25 00:16…
## $ time_label <chr> "2021-04-25 00:15:10", "2021-04-25 00:15:10", "2021-04-25 0…
We now have minute-level GPS points over multiple days.
Movement Analytics
Now we will calculate:
- Distance between consecutive points
- Time difference
- Speed
This helps us to have an idea of how fast we are moving at any point!
gps_data <- gps_data %>%
mutate(
lon2 = lead(longitude),
lat2 = lead(latitude),
t2 = lead(time),
dt_s = as.numeric(difftime(t2, time, units = "secs")),
step_m = distHaversine(
cbind(longitude, latitude),
cbind(lon2, lat2)
),
speed_mps = step_m / dt_s
) %>%
filter(dt_s > 0) %>% # remove zero/negative time gaps
mutate(speed_mps = ifelse(is.infinite(speed_mps), NA, speed_mps))
summary(gps_data$speed_mps)## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 0.0000 0.0000 0.0000 1.2290 0.8059 884.9334
That is a dynamic view of human mobility in meters per second. Some of those numbers are a little high though!
Mapping with Leaflet: Interactive Map Colored by Speed (mph)
Now let’s use the leaflet package to make an interactive map of our GPS data. This lets us explore movement patterns dynamically, and it’s a great way to get familiar with spatial data in a hands-on way. Let’s start off by converting speed to miles per hour and removing unrealistic values (>90 mph).
gps_speed_clean <- gps_data %>%
mutate(speed_mph = speed_mps * 2.23694) %>%
filter(
!is.na(speed_mph),
speed_mph > 0,
speed_mph < 90
)Now let’s map points by speed!
pal_speed <- colorNumeric(
palette = "viridis",
domain = range(gps_speed_clean$speed_mph, na.rm = TRUE)
)
leaflet(gps_speed_clean) %>%
addProviderTiles("CartoDB.Positron") %>%
addCircleMarkers(
lng = ~longitude,
lat = ~latitude,
radius = 3,
color = ~pal_speed(speed_mph),
stroke = FALSE,
fillOpacity = 0.85,
popup = ~paste(
"Time:", time_label,
"<br>Speed (mph):", round(speed_mph, 1)
)
) %>%
addLegend(
"bottomright",
pal = pal_speed,
values = ~speed_mph,
title = "Speed (mph)"
)
This map shows where movement is slow (likely stationary or walking) versus fast (driving). What do we see?
Time-of-Day Maps
Let’s compare spatial behavior across time of day. Where did I spend time during the morning? What about afternoon and evening? What about night?
gps_data <- gps_data %>%
mutate(
hour = hour(time),
tod = cut(
hour,
breaks = c(-1, 5, 11, 17, 23),
labels = c("Night", "Morning", "Afternoon", "Evening")
)
)
gps_sf <- st_as_sf(gps_data, coords = c("longitude", "latitude"), crs = 4326)
tmap_mode("plot")## ℹ tmap modes "plot" - "view"
## ℹ toggle with `tmap::ttm()`tm_shape(gps_sf) +
tm_symbols(
fill = "steelblue",
col = NA,
size = 0.2,
fill_alpha = 0.6
) +
tm_facets(by = "tod", ncol = 2, free.coords = FALSE) +
tm_title(
text = "Activity Space by Time of Day",
size = 1.2,
) +
tm_basemap()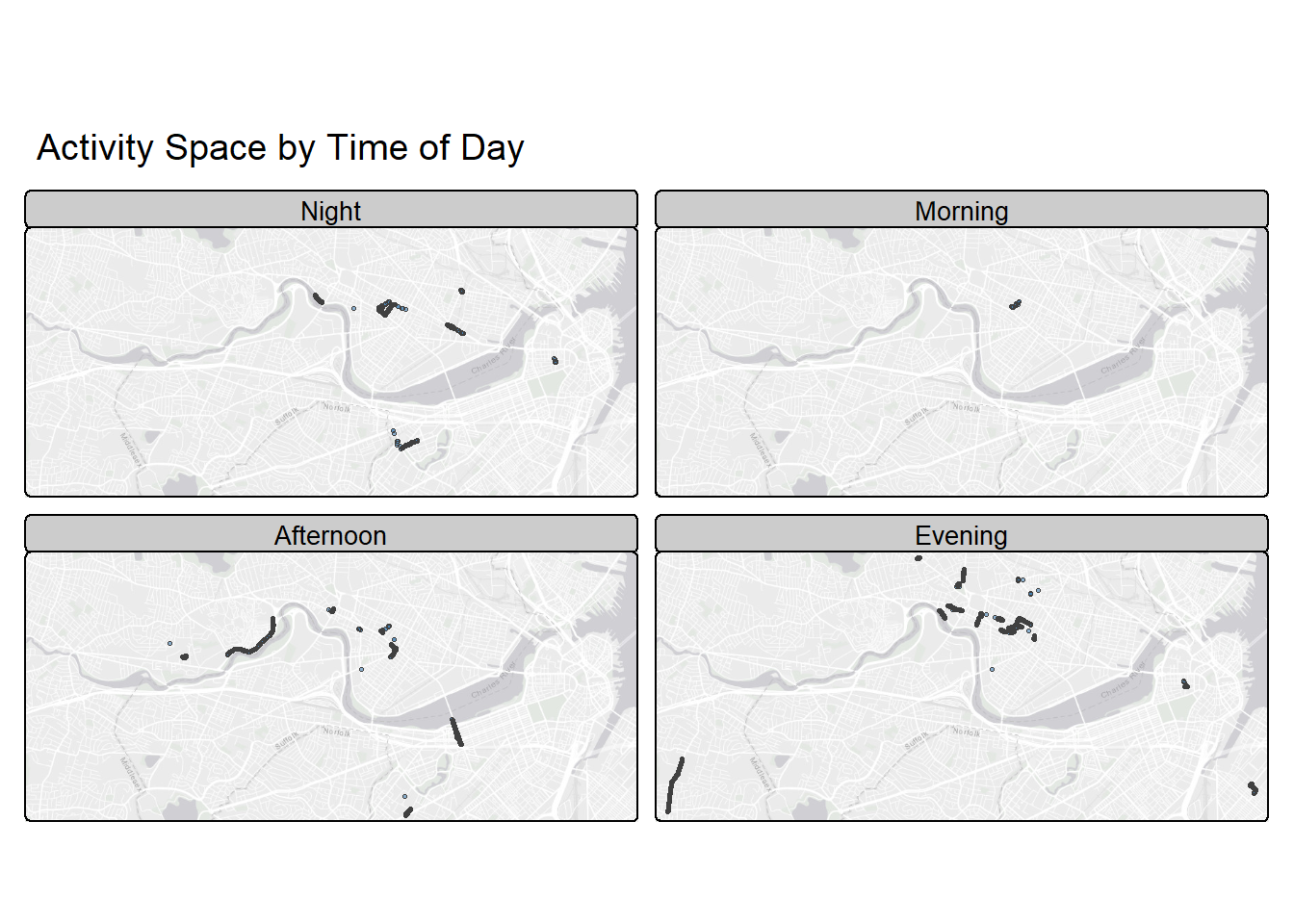
Interesting. I like the nightlife. I like to boogie.
🌿 Add Nature: Load up an NDVI Raster
Let’s make this a little richer by adding an environmental exposure that we could link to our data. Our old friend NDVI (Normalized Difference Vegetation Index) from Landsat (30m resolution) helps us understand greenness and vegetation density.
download.file("https://raw.githubusercontent.com/pjames-ucdavis/SPH215/refs/heads/main/NDVI_rast_boston.tif", destfile = "NDVI_rast_boston.tif", mode = "wb")
ndvi_raster <- rast("NDVI_rast_boston.tif")
ndvi_raster## class : SpatRaster
## size : 210, 583, 1 (nrow, ncol, nlyr)
## resolution : 0.0002694946, 0.0002694946 (x, y)
## extent : -71.19885, -71.04174, 42.33032, 42.38692 (xmin, xmax, ymin, ymax)
## coord. ref. : lon/lat WGS 84 (EPSG:4326)
## source : NDVI_rast_boston.tif
## name : NDVI_mean
## min value : -0.1340106
## max value : 0.7256226tm_shape(ndvi_raster) +
tm_raster(col.scale = tm_scale_continuous(values = "brewer.yl_gn"),
col.legend = tm_legend(title = "NDVI"))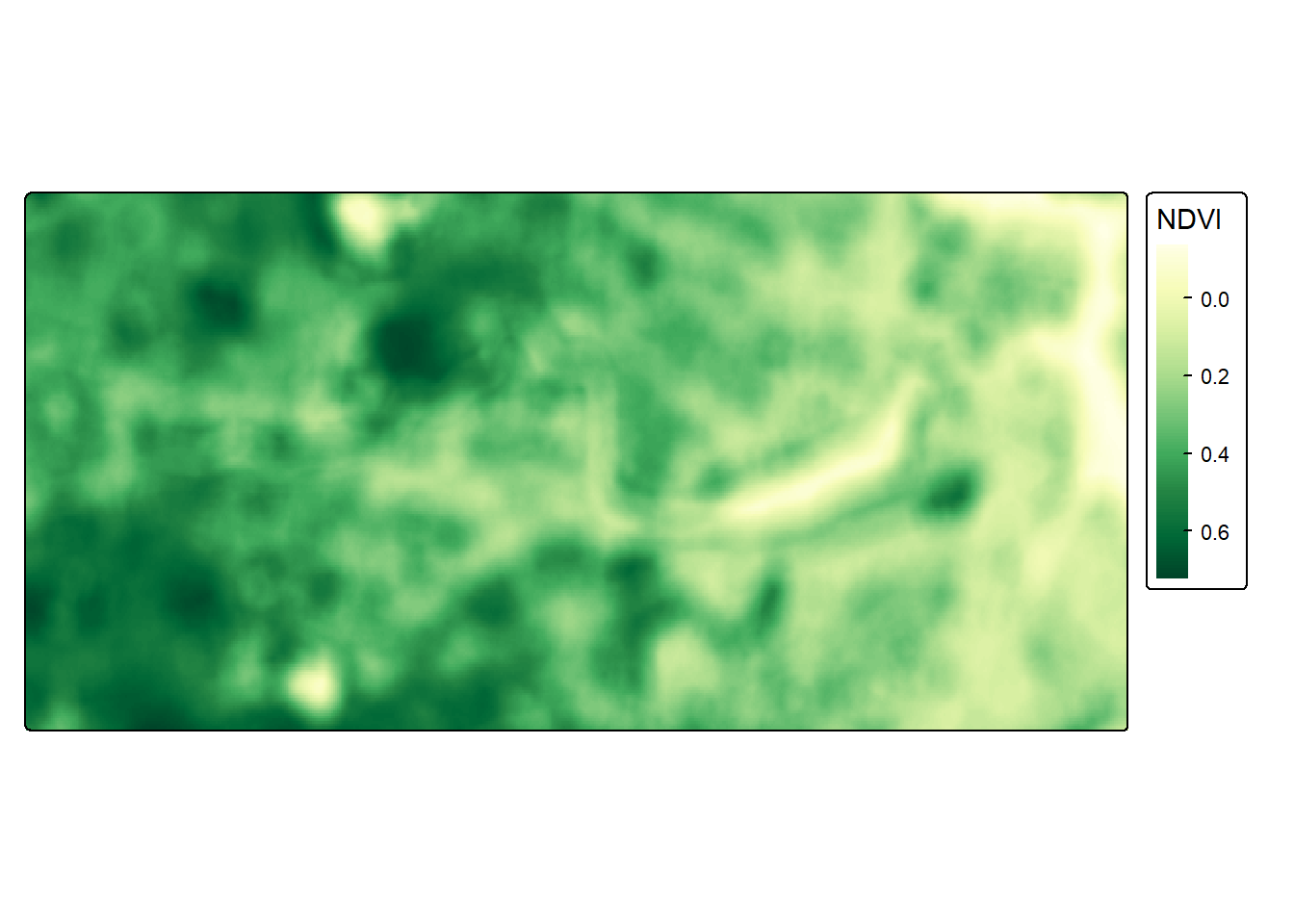
Looks quite green!
✨ Make It Beautiful with tmap
Time to show off with tmap, our old reliable thematic mapping package. We reproject the GPS points to match the raster and then we layer it all together.
tmap_mode("view") # switch to interactive map mode## ℹ tmap modes "plot" - "view"# Reproject GPS CRS to match raster
gps_sf <- st_as_sf(gps_data, coords = c("longitude", "latitude"), crs = 4326)
# Transform our GPS CRS to match the raster!
gps_proj <- st_transform(gps_sf, crs = crs(ndvi_raster))
# Map it!
tm_shape(ndvi_raster) +
tm_raster(col.scale = tm_scale_continuous(values = "brewer.yl_gn"),
col_alpha = 0.55,
col.legend = tm_legend(title = "NDVI")) +
tm_shape(gps_proj) +
tm_symbols(
fill = "blue",
size = 0.5,
col = NA,
fill_alpha = 0.1
) +
tm_title_out(title = "Where We've Been with a Little Green)")## Registered S3 method overwritten by 'jsonify':
## method from
## print.json jsonlite## Warning: tm_scale_intervals `label.style = "continuous"` implementation in view mode
## work in progress